This post has been republished via RSS; it originally appeared at: Microsoft Research.

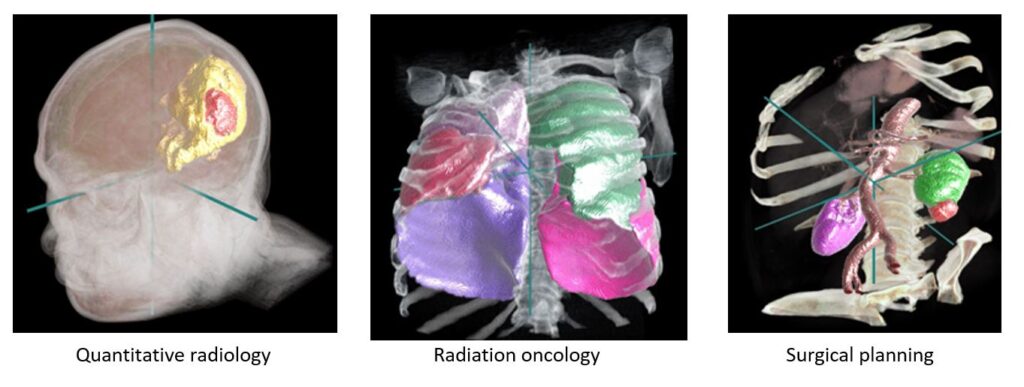
For over a decade, the Project InnerEye team at Microsoft Research Cambridge has been developing state-of-the-art machine learning methods for the automatic, quantitative analysis of three-dimensional medical images. An important application is to assist clinicians for image preparation and planning tasks for radiotherapy cancer treatment. This task involves a radiation oncologist or specialist technician manually examining and marking up dozens of 3D Computed Tomography (CT) image scans. This may take one or more hours currently, depending on the type of cancer. Our research shows that machine learning (ML) can help reduce this burden on clinicians by decreasing the time for doing this task to a few minutes.
Project InnerEye has been working closely with the University of Cambridge and Cambridge University Hospitals NHS Foundation Trust to make progress on this problem through a deep research collaboration. Dr. Raj Jena, Group Leader in machine learning and radiomics in radiotherapy at the University of Cambridge, explains, “The strongest testament to the success of the technology comes in the level of engagement with InnerEye from my busy clinical colleagues. For over 15 years, the promise of automated segmentation of images for radiotherapy planning has remained unfulfilled. With the InnerEye ML model we have trained on our data, we now observe consistent segmentation performance to a standard that matches our stringent clinical requirements for accuracy.”
The goal of Project InnerEye is to democratize AI for medical image analysis and empower developers at research institutes, hospitals, life science organizations, and healthcare providers to build their own medical imaging AI models using Microsoft Azure. So to make our research as accessible as possible, we are releasing the InnerEye Deep Learning Toolkit as open-source software. The toolkit makes it easy to train high-performance models with ensembles and deploy your models using Azure Machine Learning or Azure Stack Hub. We’re excited to see how people and organizations build on this to improve patient care.
Open-source InnerEye Deep Learning Toolkit
Developing ML models for medical imaging is advancing rapidly as new techniques, such as deep neural networks, continue to improve. It can be difficult to focus on core ML advances due to the complex software engineering and compute infrastructure needed to define, train, test, and track their projects. The Project InnerEye team has worked on a variety of ML projects (including radiotherapy, surgical planning, and quantitative radiology) and developed a common framework for our own team to streamline our workflow, taking advantage of Microsoft Azure. We are delighted to share this toolkit as open source for anyone who wants to:
- Build state of the art 3D/2D medical imaging classification, segmentation, or sequential models at scale.
- Avoid managing and maintaining an elastic cutting-edge GPU cluster.
- Follow best practices to build AI models—DevOps for ML (MLOps).
We have created a configuration-based approach for building your own image classification, segmentation, or sequential models. This uses templates for different models, including common scenarios such as radiotherapy segmentation, radiology segmentation, and ophthalmology classification. Classification, regression, and sequence models can be built with only images as inputs, or a combination of images and non-imaging data as input. This supports typical use cases on medical data where measurements, biomarkers, or patient characteristics are often available in addition to images. You can use pre-configured neural networks, such as UNet3D, or bring your own networks.

One of the challenges of building state-of-the-art models is training on modern compute architectures at scale. Traditionally this involves building and maintaining elastic clusters of GPUs, which is not what ML researchers and developers necessarily want to spend their time doing. Cloud computing makes it easier to manage distributed training across large numbers of GPUs. The InnerEye Deep Learning Toolkit takes advantage of Azure Machine Learning Services for:
- Scaling clusters to many compute nodes, to train models on multiple GPUs
- Only paying per experiment
- Saving costs by leveraging low priority nodes
- Using the latest GPUs, Intelligent Processing Units (IPUs), and Field Programmable Gate Arrays (FPGAs)
- Using advanced capabilities such as Azure Confidential Computing
To build trust in ML models, it is important to follow industry and community best practices for reliability, accountability, and transparency. Our toolkit uses Azure Machine Learning to manage DevOps for ML (MLOps), including:
- Experiment traceability as Azure Machine Learning keeps a full record of all experiments that were executed, including a snapshot of the code. Tags are added to the experiments automatically, that can later help filter and find old experiments.
- Experiment transparency so all team members have access to each other’s experiments and results.
- Model reproducibility, for example two model trainings run using the same code and data will result in the same metrics. All sources of randomness, like multithreading, are controlled for.
- Model management, including versioning models with rich metadata.
- Model deployment using Azure Machine Learning, or Azure Stack Hub as part of an Intelligent Edge solution.
- Integration with Git and Continuous Integration (CI).
In addition, the toolkit supports more advanced ML development features including:
- Cross-validation using Azure Machine Learning’s built-in support, where the models for individual folds are trained in parallel. This is particularly important for the long-running training jobs often seen with medical images.
- Hyperparameter tuning using HyperDrive with Azure Machine Learning.
- Building ensemble models by combining multiple diverse models to improve predictive performance
- Comparing new models against a baseline model by using statistical tests
- Creating new models easily via a configuration-based approach and inheriting from an existing architecture.
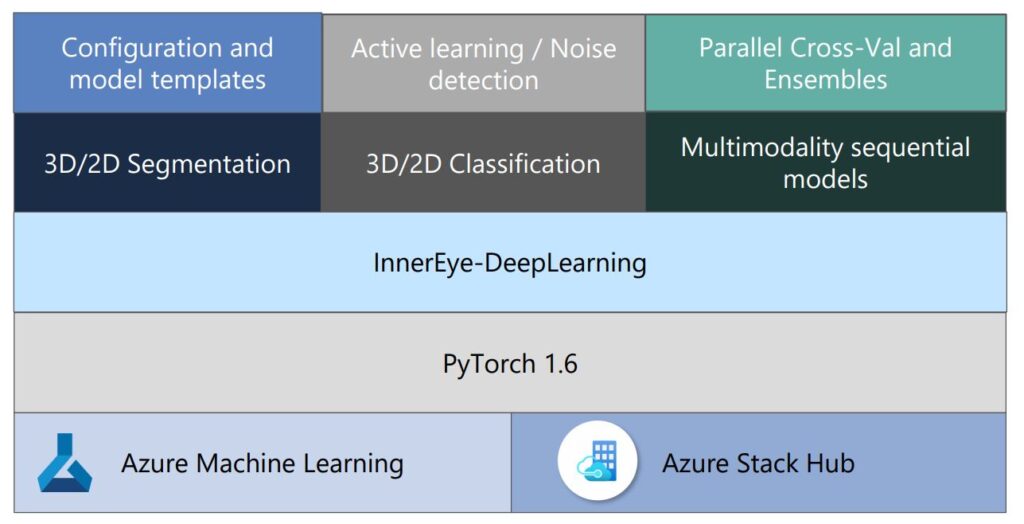
The InnerEye Deep Learning Toolkit has been designed with usability and flexibility at its core, built on PyTorch and making extensive use of Microsoft Azure.
Toward clinical use of state-of-the-art machine learning
The Topol Review of the NHS emphasized that technology should give clinicians “the gift of time.” Our InnerEye technology aims to provide the building blocks to allow users to do exactly this.
We have released the InnerEye Deep Learning Toolkit as open-source software on GitHub to make this ML library and technical components available to as many people and organizations as possible. The InnerEye Deep Learning Toolkit can be used by researchers to build and refine their own models and apply them in many ways, including applications yet to be thought of. Healthcare providers, companies, and partners may use this toolkit to develop their own ML products and services, including Azure Stack Hub (subject to testing and regulatory approval as appropriate, such as FDA clearance, CE marking, or in-house exemption controls). Azure Stack Hub extends Azure services and capabilities to customers’ environment of choice, from datacenters to the edge. Customers can build, deploy, and run hybrid and edge apps consistently across their IT ecosystem and take advantage of cloud native services such as AI and ML. This extension of Azure allows customers to comply with regulatory requirements and enables latency-sensitive workloads.
Dr. Jena adds, “With the potential to refine and take ownership of the models ourselves through technology like Azure Stack Hub, we see a way that we can integrate machine learning technologies into our treatment pathway as a long-term solution that can grow and evolve over time.”
We are already supporting several research teams in using the InnerEye Deep Learning Toolkit to build their own ML models. We invite anyone interested in medical imaging AI to join and contribute to the open-source project. We welcome all contributions, no matter how small, whether using the toolkit, filing issues and bugs, or writing and extending the toolkit in new directions. We look forward to the research and healthcare technology community building on this foundation to ultimately benefit patients around the world.
Acknowledgments
Project InnerEye is the work of a multi-disciplinary team, including Javier Alvarez-Valle, Junaid Bajwa, Shruthi Bannur, Melanie Bernhardt, Melissa Bristow, David Carter, Ben Glocker, Pratik Ghosh, Raj Jena, Usman Munir, Hannah Murfet, Jay Nanavati, Aditya Nori, Kenton O’Hara, Ozan Oktay, Nadzeya Paleyes, Anton Schwaighofer, and Kenji Takeda. We are grateful for the deep partnerships with Ivan Tarapov, Michela Sainato, Alex Sutton, and our colleagues across Microsoft Research, Microsoft Healthcare, and Microsoft Azure. We sincerely thank all our external collaborators and early adopters for their feedback and contributions.
Disclaimer: The InnerEye Deep Learning Toolkit (the “Toolkit”) is a research tool provided AS-IS for use by third parties in their design and testing of machine learning models. This Toolkit is not intended or made available for clinical use as a medical device, clinical support, diagnostic tool, or other technology intended to be used in the diagnosis, cure, mitigation, treatment, or prevention of disease or other conditions. This Toolkit is not designed or intended to be a substitute for professional medical advice, diagnosis, treatment, or judgment and should not be used as such. All users are responsible for reviewing the output of the developed model to determine whether the model meets the user’s needs and for validating and evaluating the model before any clinical use. Microsoft does not warrant that the Toolkit or any materials provided in connection therewith will be sufficient for any medical purposes or meet the health or medical requirements of any person.
The post Project InnerEye open-source deep learning toolkit: Democratizing medical imaging AI appeared first on Microsoft Research.
