This post has been republished via RSS; it originally appeared at: New blog articles in Microsoft Community Hub.
Azure Machine Learning is a service that enables you to create, train, deploy, and manage machine learning models and experiments. You can also use it to run HPC applications, such as simulations, rendering, or data analysis.
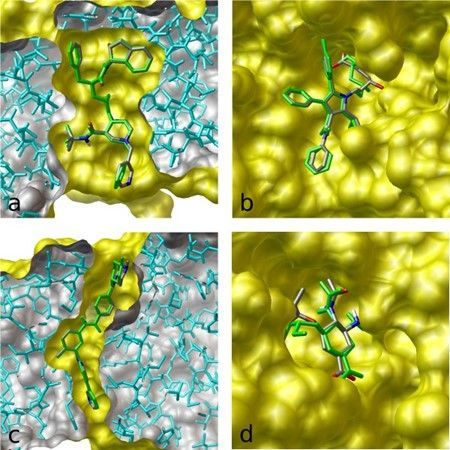
Autodock is a powerful tool for molecular simulation, essential for researchers who work with large-scale computational tasks in HPC environments. It is especially useful for drug design and biomolecular interaction studies.
This article will guide you through the process of running Autodock Vina docking simulation scripts on the Azure Machine Learning platform. You will also learn how to provision Azure NetApp Files (ANF) volumes for the Machine Learning notebooks, which can provide persistent storage for most HPC applications.
The steps are:
- Prepare your Azure Machine Learning environment
- Create ANF volume
- Install Autodock Vina on the ANF volume
- Create a compute cluster to run your job
- Create your Autodock Vina docking script
- Create your command job to run the docking script on the compute cluster
At the end, we will conclude this tutorial and highlight the benefits of using AML environment for HPC applications. We will also explore some customization options based on various needs and factors.
Let’s begin!
- Prepare your Azure Machine Learning environment
- Follow Quickstart: Get started with Azure Machine Learning to create an Azure Machine Learning workplace, sign in to studio and create a new notebook.
- Set your kernel and create handle to workplace. You'll create ml_client for a handle to the workspace. Enter your Subscription ID, Resource Group name and Workspace name and run in your notebook.
- Create ANF volume
- Follow Create an NFS volume for Azure NetApp Files with Service Level “Standard” and NFSv3 protocol to create a 4TiB size volume.
To ensure accessibility from Azure Machine Learning compute to the ANF volume, we will place both the Azure Machine Learning compute resources and the Azure NetApp Files subnets within the same VNET.
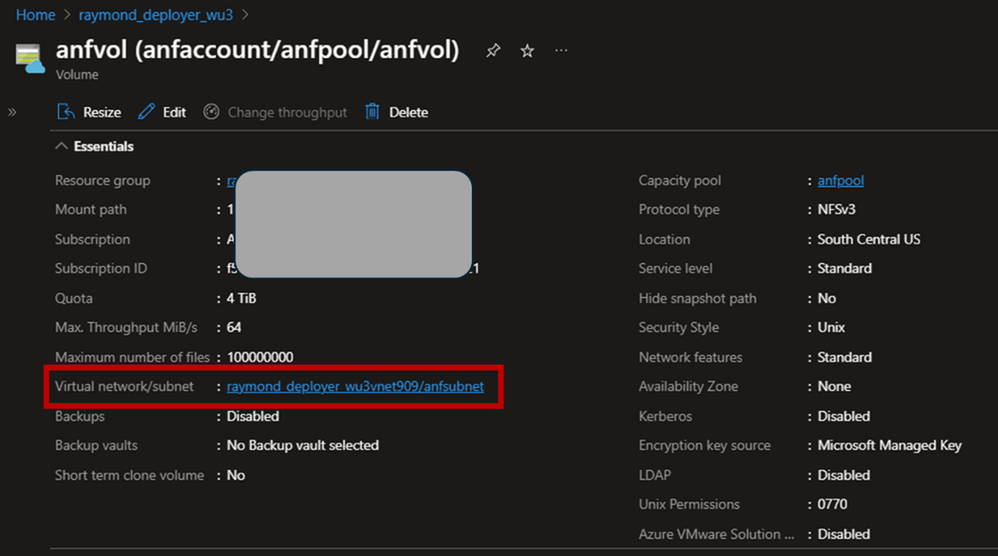
2. Open terminal of your notebook. Follow Mount NFS volumes for virtual machines | Microsoft Learn to test if you can mount the ANF volume from the Compute node successfully.
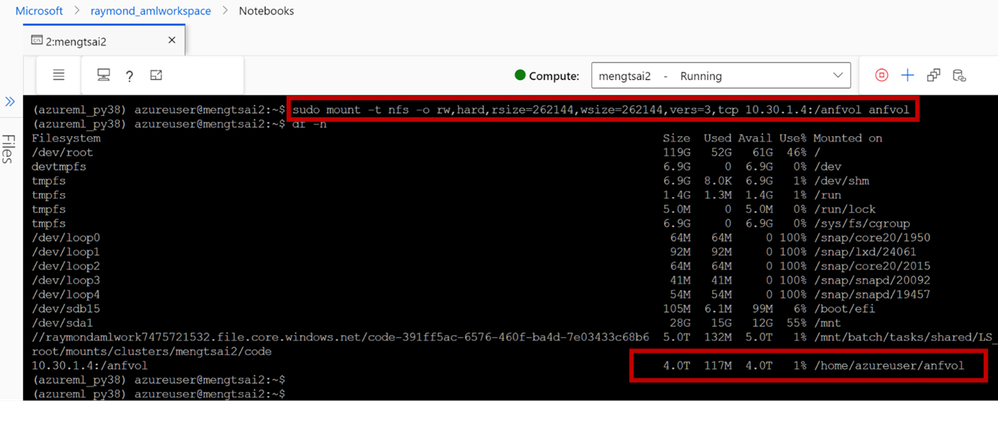
- Install Autodock Vina on the ANF volume
Follow Installation — Autodock Vina 1.2.0 documentation (Autodock-vina.readthedocs.io) to install Autodock Vina. We will download the executable for the latest release (1.2.5):
- Create a compute cluster to run your job
- Create your Autodock Vina docking script (basicdocking.sh)
In this example, we will dock the approved anticancer drug imatinib (Gleevec; PDB entry 1iep) in the structure of c-Abl using AutoDock Vina. The target for this protocol is the kinase domain of the proto-oncogene tyrosine protein kinase c-Abl. The protein is an important target for cancer chemotherapy—in particular, the treatment of chronic myelogenous leukemia. We set exhaustiveness value to 32 in order to get a more consistent docking result.
This script will run AutoDock Vina to perform molecular docking and generate a PDBQT file named 1iep_ligand_ad4_out.pdbqt. This file will include all the poses that the script discovered. The script will also display docking information on the terminal window.
- Create your command job to run the docking script on the compute cluster
- Submit your job
The file 1iep_ligand_ad4_out.pdbqt or the terminal window will display results similar to the ones below after the job has run successfully. The predicted free energy of binding is approximately -13.22 kcal/mol, as shown.
Summary and discussion
- As illustrated in this article, you can use the Azure Machine Learning studio, a web interface that provides a user-friendly and collaborative environment for building and managing your machine learning and HPC projects. You can also use the Azure Machine Learning SDKs and CLI to programmatically interact with the platform and automate your tasks.
- You can utilize the scalability and flexibility of Azure to run your HPC applications on demand, without investing in costly and complex on-premises infrastructure. You can pick from various VM sizes and configurations that fit your needs, including the latest NVIDIA and AMD GPUs with high-bandwidth InfiniBand interconnect.
- Azure Machine Learning pricing is transparent and easier for customers to estimate cost. The pricing is based on the amount of compute used, and there are no additional charges for using Azure Machine Learning. You can pay for compute capacity by the second, with no long-term commitments or upfront payments. You can also save money across select compute services globally by committing to spend a fixed hourly amount for 1 or 3 years, unlocking lower prices until you reach your hourly commitment. Azure also provides a pricing calculator that allows you to estimate the cost of using Azure Machine Learning running your HPC applications.
- Storage options. In our example, we have implemented Azure NetApp Files (ANF) as the persistent storage solution. Azure offers a range of alternative storage options tailored to specific performance requirements such as IOPS, throughput, and cost considerations. Including Azure Managed Lustre Storage, Azure Files, and Azure Blob, each serving distinct needs based on your application's demands and budget constraints.
Please note that Azure Machine Learning is not inherently designed for running HPC applications. Therefore, you may have to modify or rewrite some parts of your code during the migration process. For example, you may need to create a parallel pipeline or manually handle the logic for distributing the workload among nodes.
