This post has been republished via RSS; it originally appeared at: New blog articles in Microsoft Tech Community.
A GWA (Genome-wide association) study is an observational study of a genome-wide set of genetic variants in different individuals to see if any variant is associated with a trait. A GWA study typically focuses on associations between single-nucleotide polymorphisms (SNPs) and traits like major human diseases, but can equally be applied to identify trait-associated variants in other organisms, including animals, plants, viruses, and bacteria. SNPs are single nucleotide (A,G,C,T) variants that occur across a population at specific genomic positions, and can be associated with phenotypic traits of interest.
With the increasing availability and affordability of genome-wide sequencing and genotyping technologies, biomedical researchers are faced with increasing computational challenges in managing and analyzing large quantities of genetic data. Previously, this data intensive research required computing and personnel resources accessible only to large institutions. Today, mature and cost-effective Cloud computing allows researchers to analyze their data without the need to acquire and maintain large, costly, local computing infrastructure, providing the following advantages:
- A secure and compliant working environment for all researchers to work together remotely with proper privilege on sensitive data.
- Various CPU/GPU options and cost-effective spot instances for researchers to choose on-demand, to fulfill different computational, storage and data transfer requirements.
- Scalability of High-Performance Computing (HPC) clusters to expand research potential, prevent high upfront infrastructure investments and long-term maintenance costs.
Objective
In this article, we will examine how Azure can accelerate the workflow of a recent GWA study by National Taiwan University Medical Center. This project is aiming to establish a COVID-19 mortality prediction model leveraging UK Biobank data. There are many researchers involved and working remotely on this project. We will first introduce a practical architecture enabling all researchers to work together efficiently on Azure.
In a typical GWA study, “Quality Control” and “Genotype Imputation” are two of the most computationally intense and time-consuming phases and it’s no exceptional for this project. We will examine the performance bottleneck and suggest the most appropriate Virtual Machine (VM) to shorten the Quality Control process, which would reduce the total process time from days to minutes. We will then examine the need for scalability to support Genotype Imputation and suggest an auto-scale Azure HPC cluster approach to compute on millions of SNPs, reducing the execution time from months to days. Finally, we will walk through how to estimate total cost when doing a GWA study on Azure.
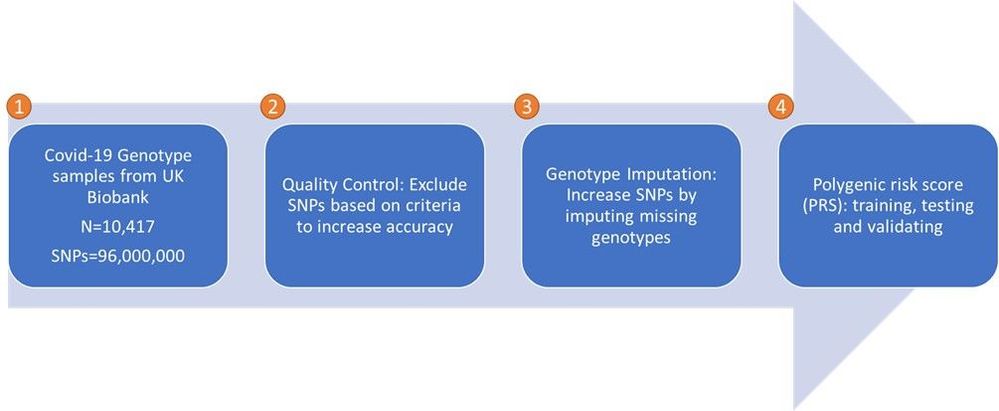
Below is the workflow of the GWA study.
Figure: The GWA workflow aiming to establish a COVID-19 mortality prediction model leveraging UK Biobank data
- Covid-19 genotype samples are fetched from UK Biobank.
- There are 10,417 Covid-19 samples as of May 2021, and each individual sample consists of around 96 million of SNPs.
- Those samples consist of 22 chromosome files (minus the sex chromosome, #23). The file size spans from the smallest at 37 GB (chromosome #22) to the largest at 188 GB (chromosome #2). To total size is ~2.4 TB.
- Quality Control is an extremely important step in GWA studies, as it can dramatically reduce the number of false positive associations. PLINK is considered the most popular tool in the field to do the quality control. We will examine how to improve performance on Azure VMs with NVMe disks when running Quality Control using PLINK.
- Genotype Imputation generates missing genotypes to improve data quality, which in turn increases the power and resolution of genetic association studies. Beagle, IMPUTE, MACH, Minimac, and SHAPEIT are among the most popular and cited tools in this area. All those imputation tools are memory-bound and use only CPU, no GPU. Among those tools, Beagle has demonstrated better performance and lower memory usage while maintaining similar imputation accuracy [ref1, ref2, ref3]. We will examine how to scale the imputation jobs on an HPC cluster in a cost-effective approach.
- Polygenic risk score: PRS gives an estimate of how likely an individual is to have a given trait related to genetics. We will not examine this phase in this article.
Architecture and GWA study workflow on Azure
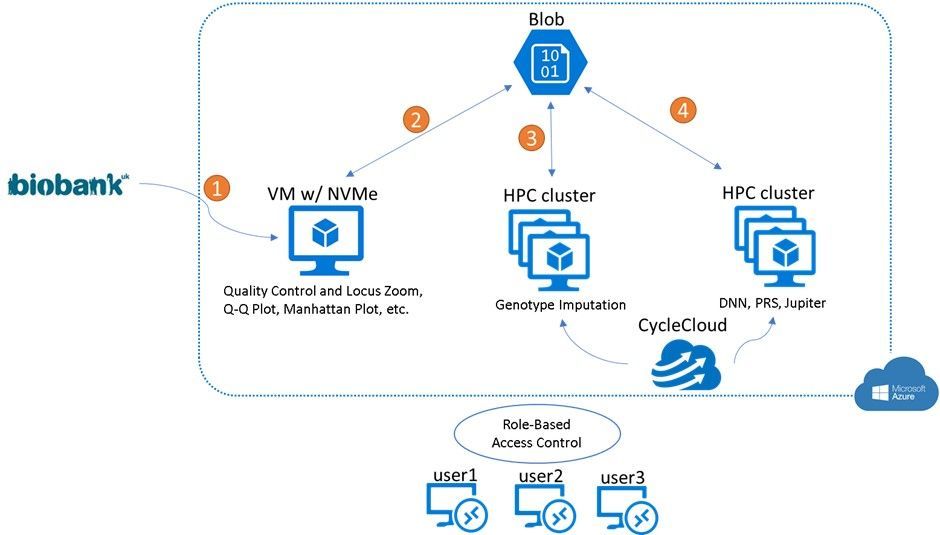
Below is the architecture and workflow researchers can use to work remotely on this GWA study on Azure.
Table: The Azure architecture and workflow when working on the GWA study
Security
Users access the Azure resource to manipulate data with credential-protected access privileges. Azure RBAC (Role-Based Access Control) is one of the key features to achieve this goal.
Fast Network Transfer
Users will enjoy the up to 32 Gbps network bandwidth when transferring files among different VMs. It takes less than 20 minutes to transfer 2.4 TB raw data from Azure Blob to the VMs.
Compute Performance
Azure VMs with local NVMe disks are suggested to execute quality control as well as huge data manipulation, including fetching dataset, genotype files conversion, and files transmission. NVMe disks’ superfast IO can minimize CPU’s waiting time for data, as well as to maximize the network transmission throughput.
High Performance Computing Cluster
HPC cluster is managed by Azure CycleCloud, which can provision necessary compute VMs in seconds for any intensive compute purpose. And shut down those VMs immediately when the job is completed to save the overall cost.
Data Management
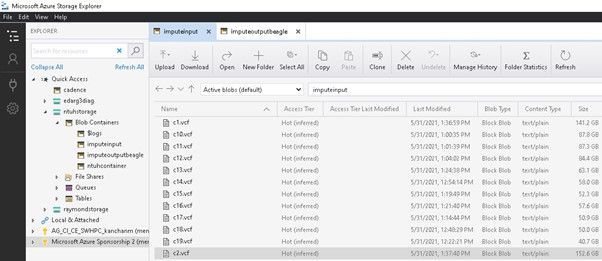
A typical GWA study can involve many data manipulation, conversion, and archiving steps, which requires an inexpensive storage solution for intermediate data. Azure Blob is suitable for this requirement. Users will also find the GUI interface of Azure Storage Explorer useful when managing multiple folders and versions of files on different phases of the workflow.
Figure: Azure Storage Explorer provides intuitive and feature-rich graphical user interface (GUI) for full management of Azure storage resources
Improve Quality Control performance by using Azure VMs with NVMe disks
The need for careful attention to data quality has been appreciated in the GWA field. PLINK is the most popular tool to perform genotype quality control. PLINK is a free, open-source whole genome association analysis toolset, designed to perform a range of basic, large-scale analyses in a computationally efficient manner. Below is the command we examined.
What this command does is to first read a file “chr2.bgen“ as input. It is a bgen format (Oxford Genotype) file of chromosome #2, which contains around 7.1 million SNPs with 10,417 samples. The size of this file is 188 GB. This command will then exclude SNPs based on specified criteria to increase genotype quality, exporting results as another file, similar in size to the input file. This means it require at least double size of the input files on your working disk.
The table below shows the results of running the quality control job on a number of different VMs, along with CPU, memory, local disk, and price information. Among them, HC44rs, HB120v3 and L80sv2 have NVMe disks, and E64dsv4 does not. For HB120v3 and L80sv2, we striped the NVMe disks to one single RAID 0 array for best I/O performance, which becomes 1.92 TB and 19.2 TB respectively. And we regard the local disk as working disk which will store both input and output files.
Table: CPU, Memory, Disks and Price of Virtual Machines we examined
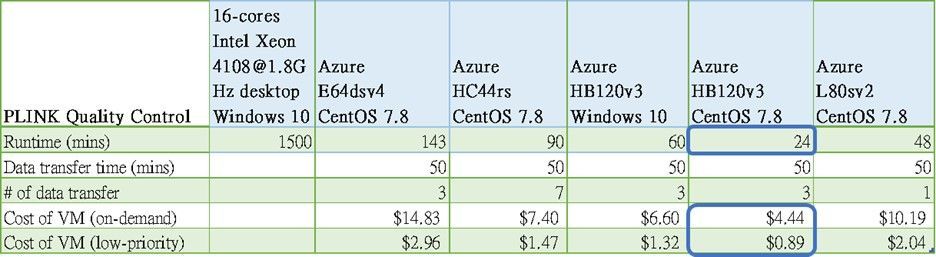
The table below shows the QC runtime, data transfer time and VM cost estimation. As you can see this PLINK job took over 24 hours to complete on an on-premises 16-core Intel Xeon 4108@1.8GHz Windows 10 desktop. And it can be done in 24 minutes on HB120v3 with CentOS 7.8, which is 6x and 3.75x faster than E64dsv4, and HC44rs, respectively. We also observed that this command runs 2x faster on Linux than on a Windows 10 machine.
Table: PLINK Quality Control runtime, data transfer time and total cost of VM
We found disk I/O is the 1st performance bottleneck of the Quality Control task. In fact, most PLINK jobs consist of massive, small chunks of read and write operations. That’s why the overall performance will improve when you store the input and output files on the NVMe disk as it provides superfast IOPS and disk throughput.
The 2nd bottleneck will then be CPU execution and memory bandwidth. As PLINK will, by default, utilize all available CPUs, it gives HB120v3 the performance edge over other VMs as it contains 120 CPUs and has higher L3 cache bandwidth.
Please note “Data transfer time” is also considered in the above table. We would need to transfer total of 4.8 TB files in the QC phase, as the total input data is ~2.4 TB and output files size is about the same as input files. It’s estimated to take 40~50 minutes to transfer all files between Azure Blob and the striped NVMe disk, utilizing the up to 32 Gbps of Azure network backbone.
We also estimated the minimum “# of data transfer” efforts. For example, L80sv2 contains a total of 19.2 TB NVMe RAID 0 disk, which is big enough for you to do only 1 iteration of download and upload effort to transfer all input and output files. You can download 2.4 TB input files in the disk, execute the QC jobs, and upload back the 2.4 TB output files to the Blob storage. In this case, it provides researchers the “jobs-packing” option to fully utilize the large number of CPUs of a VM. That is, you can run multiple or even all chromosomes’ QC jobs simultaneously in one VM at one time. Which will maximize the overall throughput, minimize data transfer efforts, and save cost.
Scale the Genotype Imputation jobs on HPC cluster
Genotype Imputation is now an essential approach in the analysis of GWA studies. The technique is to accurately evaluate the evidence for association at genetic markers that are not directly present in the genetic sequence. Recall that all popular genotype imputation techniques are memory-bound tasks. And Beagle is one of the most popular tools for Genotype Imputation, which has demonstrated better performance and lower memory usage while maintaining imputation accuracy similar to the less performant tools.
We examined Beagle’s scalability and performance by running the command below on HB120v3 with different number of threads (CPUs). Beagle is run using Java. We used the 5.2-29May21.d6d version, which was released on May 21, 2021.
The greater the number of SNPs and samples in the input file, the more memory is needed to hold the working data. We specify 400 GB (-Xmx400g) as the upper bound of the memory pool to use as much as possible of the 448 GB memory of HB120v3 VM. We found it requires this size of memory to impute one and only one single chromosome file on one VM. The “OutOfMemoryError” error will be thrown if not specifying enough heap memory. For scenarios needing even more memory, you can consider the M192idmsv2 VM, which comes with ~9x the memory capacity (4 TB) of the HB120v3.
The “gt” parameter specifies the chromosome VCF (Variant Call Format) file as input. The “window” parameter controls the amount of computer memory that is used, and 10 is the minimal suggested number. And the “nthreads” specifies the number of threads (CPUs) to be used.
We examined 2 chromosome files with different number of SNPs:
- c9.vcf (chromosome #9):
- ~2.1 million SNPs
- Input: 78 GB
- Output: 15 GB
- c21.vcf (chromosome #21):
- ~0.632 million SNPs
- Input: 25 GB
- Output: 0.5 GB
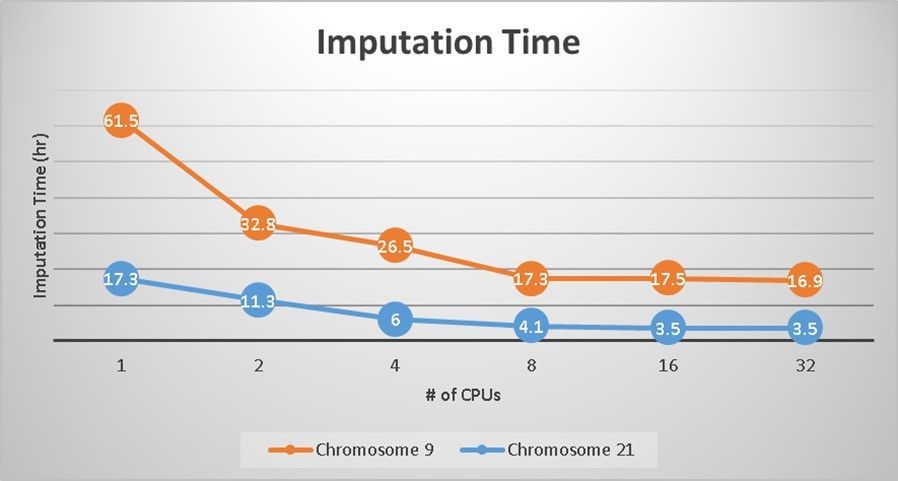
Below the results of imputation time (hr) of both chromosomes, running on 1, 2, 4, 8, 16 and 32 CPUs. You can see both chromosomes scale well until 8 CPUs, and then stay flat for 16 and 32 CPUs.
Imputation performance can be affected by many factors, in which number of SNPs and number of samples the most important ones. Since we have the same 10,417 sample size of all chromosomes, we can then examine how number of SNPs affect the imputation performance.
Chromosome #9 has ~3x (2.1/0.632) more SNPs than chromosome #21, and it’s running ~5x (17.5/3.5) slower when running on 16 CPUs. Based on this information we can further estimate the imputation runtime of other chromosomes by extrapolation later.
Chart: Imputation time (hr) of #9 and #21 chromosomes on 1, 2, 4, 8, 16 and 32 CPUs
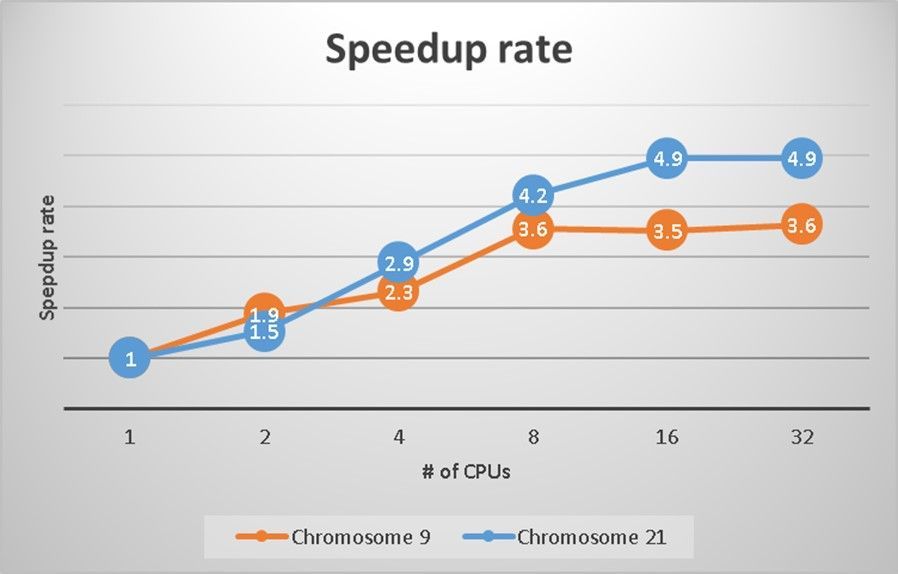
Below charts shows the speedup rate while adding more CPUs. You can see 8 CPUs the suggested sweet spot to run imputation, it will gain ~4x better performance compared to 1 CPU.
Chart: Speedup rate of #9 and #21 chromosomes on 1, 2, 4, 8, 16 and 32 CPUs
Azure CycleCloud for running Genotype Imputation on HPC Cluster
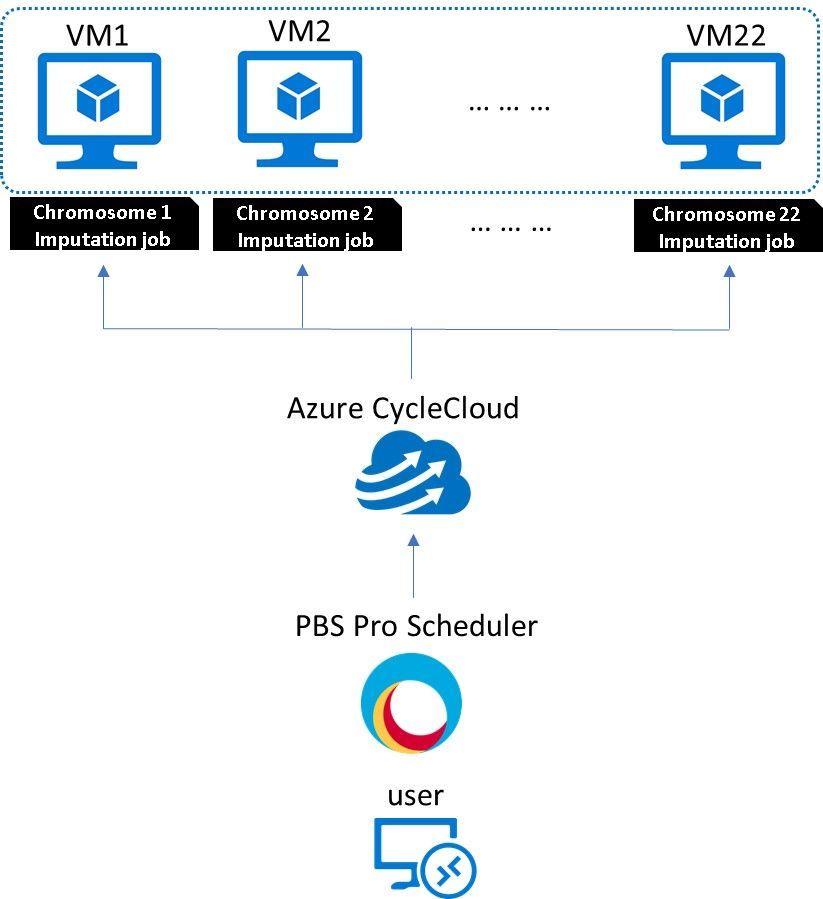
An Azure HPC cluster can perform Genotype Imputation on all 22 chromosomes simultaneously, leveraging Azure CycleCloud to parallelize the pipeline execution.
Figure: Azure HPC cluster for Genotype Imputation on all 22 chromosomes leveraging Azure CycleCloud to parallelize the pipeline execution
The User submits imputation jobs for the 22 chromosomes files through PBS Pro job scheduler, and use “place=scatter:excl” parameter to run each job exclusively in one VM.
Azure CycleCloud will then provision 22 VMs (HB120v3 in this case) in seconds, distribute jobs to each VM, run the imputation jobs in parallel, and shut down the VMs when completed.
That is, the imputation time to complete all 22 chromosomes will be determined by the largest chromosome, which is chromosome #2, as it contains the greatest number of SNPs and would take the longest time to complete. We estimated the chromosome #2 can be completed in 48 hours, based on the extrapolation of chromosome #9 and #21’s SNPs number.
Based on the same estimation methodology, the total accumulated compute time to complete all 22 chromosomes would be around 400 hours.
Please note above suggested architecture assumes that only one job running on one VM, this restriction is due to the high memory needs of the huge number of SNPs dataset. For smaller dataset, it is suggested to run multiple jobs in one VM (“jobs-packing”) when possible, so to save overall cost and still enjoy high throughput of the HPC cluster.
Computational cost of Genotype Imputation
Let’s further estimate the total cost to complete Genotype Imputation for all 22 chromosomes on Azure. The estimation will be based on below cost structure, including Compute, Storage and Networking. And the same cost estimation methodology can also be applied to other genomics analysis projects running on Azure.
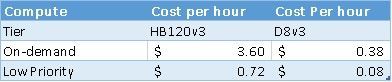
- Compute:
- HB120v3 as the compute node, D8v3 as the PBS Pro and CycleCloud master node. Both “On-demand” and “Low Priority” prices are listed.
- Storage:
- 1-year reserved price.
- P40 (2TB) for hot-tier storage, Blob (10TB) for intermediate data and archive.
- Bandwidth:
- 1-year reserved price for 10TB bandwidth.
Adding up all together, the cost to complete Genotype Imputation for all 22 chromosomes of the 10,417 samples using Beagle 5.2 would be ~$3,600 for on-demand price, which is ~$0.35 per sample. If using Low-Priority VMs, the total cost will be reduced to ~$2,324, which is ~$0.22 per sample.
Summary
In this article, we examined how a genome-wide association study (GWAS) can be executed efficiently and cost-effectively on Azure. We used a real-world Covid-19 mortality research workflow and UK Biobank dataset as our example, which consists of 10,417 samples with ~96 million of SNPs. We suggested a secure working environment on Azure for all researchers to work remotely with role-privileged control. We explored two of the most computational-intensive and time-consuming scenarios: “Quality Control” and “Genotype Imputation”, and provided performance, scalability, and cost-saving suggestions.
Regarding Quality Control using PLINK, we found Azure VMs with NVMe disks perform the best on the execution. For example, HB120v3 comes with 1.92 TB NVMe disks and 120 CPUs, or L80sv2 with 19.2 TB NVMe disks and 80 CPUs. We also suggest the “jobs-packing” strategy to maximize the overall throughput, minimize data transfer efforts, and save cost.
As for the Genotype Imputation, which is arguably the most computational and time-consuming job on GWA study, especially when the dataset contains a huge number of SNPs and samples. We examined the scalability by running different number of CPUs on HB120v3, and then suggested a scalable HPC cluster environment to impute all 22 chromosomes. Finally, we estimated the cost of performing genotype imputation jobs on Azure, which can be lower as $0.22 per sample.
Along the way, we also examined several key points on how Azure is a great genomic analytics platform for genomics researchers and medical organizations.
- Researchers can work with their familiar tools and pipelines on Azure, either on Windows or Linux machines.
- Azure provides you with high memory and HPC-class CPUs and GPUs virtual machines to help you get results efficiently.
- Azure CycleCloud help you scale up and down HPC clusters so to pay only what you use to reduce cost.
- Azure Blob as a secure and cost-effective storage by providing role-based access control (RBAC) with encryption at rest feature.
- Azure Storage Explore with intuitive GUI for data versioning, transferring and management.
- Azure’s 32 Gbps network throughput allow you to transfer TBs of data in minutes.
- Price predictability of your GWA and any other genomics projects.